|
|
| Alignments download: |
|
|
|
The alignments of 211 introns belonging to IE major subgroup, from Li and Zhang (2005), Nucleic Acids Res., 33, 2118-2128, could be downloaded HERE.
|
| .sto files: in stockholm format, alignments used by INFERNAL package. |
|
| sali alignment format: |
The Group I intron structural alignment format, ".sali", is
defined as below:
- Each entry has its sequence and structure, which means two lines;
- Non-pseudoknot base-pairings are marked by '<' and '>';
- Non-paired bases are marked by '.';
- For the two pseudoknots, P3-P7 and P1-P10, P7 and P10 are marked by '[' and ']';
- Gaps in the sequence and structure are both marked by '-';
- The insertion sequences are marked by '(number of nt)' in the sequence, and by '~' in the structure;
- The base which is competed by two different bases in P10 and P1' respectively, is marked by '^'.
|
| e.g. Figure |
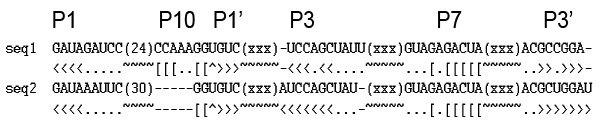
|
This format was designed to keep as much structure information in the alignment. The pseudoknot structure, overlapped basepairs are all kept. Each sequence keeps its own specific structure, although the sequences in the alignment have a consensus, which could be inferred from the alignment. For example, we have converted these alignments to be in Stockholm format, which is the accpeted format of INFERNAL package. In the process of conversion, repeat or nearly identical sequences were eliminated, and long insertions in a small part of the sequences were not kept.
|
|
|























